library(tidyverse) # general use
library(janitor) # cleaning data frames
library(here) # file/folder organization
library(readxl) # reading .xlsx files
library(ggeffects) # generating model predictions
library(gtsummary) # generating summary tables for models
# abalone data from Hamilton et al. 2022
abalone <- read_xlsx(here("data", "Abalone IMTA_growth and pH.xlsx"))Linear regression: Abalone growth and pH example
Set up
Abalone example
Data from Hamilton et al. 2022. “Integrated multi-trophic aquaculture mitigates the effects of ocean acidification: Seaweeds raise system pH and improve growth of juvenile abalone.” https://doi.org/10.1016/j.aquaculture.2022.738571
a. Questions and hypotheses
Question: How does pH predict abalone growth (measured in change in shell surface area per day, mm-2 d-1)?
H0: pH does not predict abalone growth (change in shell surface area per day, mm2d-1)
HA: pH does predict abalone growth (change in shell surface area per day, mm2d-1)
b. Cleaning
#made new data frame from abalone
abalone_clean <- abalone |>
#cleaned names with underscores and no caps
clean_names() |>
#selected for columns of interest
select(mean_p_h, change_in_area_mm2_d_1_25) |>
#changed the column names
rename(mean_ph = mean_p_h,
change_in_area = change_in_area_mm2_d_1_25)Don’t forget to look at your data! Use View(abalone_clean) in the Console.
c. Exploratory data visualization
#base layer: ggplot
ggplot(data = abalone_clean, #data is abalone_clean
aes(x = mean_ph, #x is mean_pH (predictor variable)
y = change_in_area)) + #y is change in area (response variable)
#
geom_point(size = 4,
stroke = 1,
fill = "firebrick4",
shape = 21)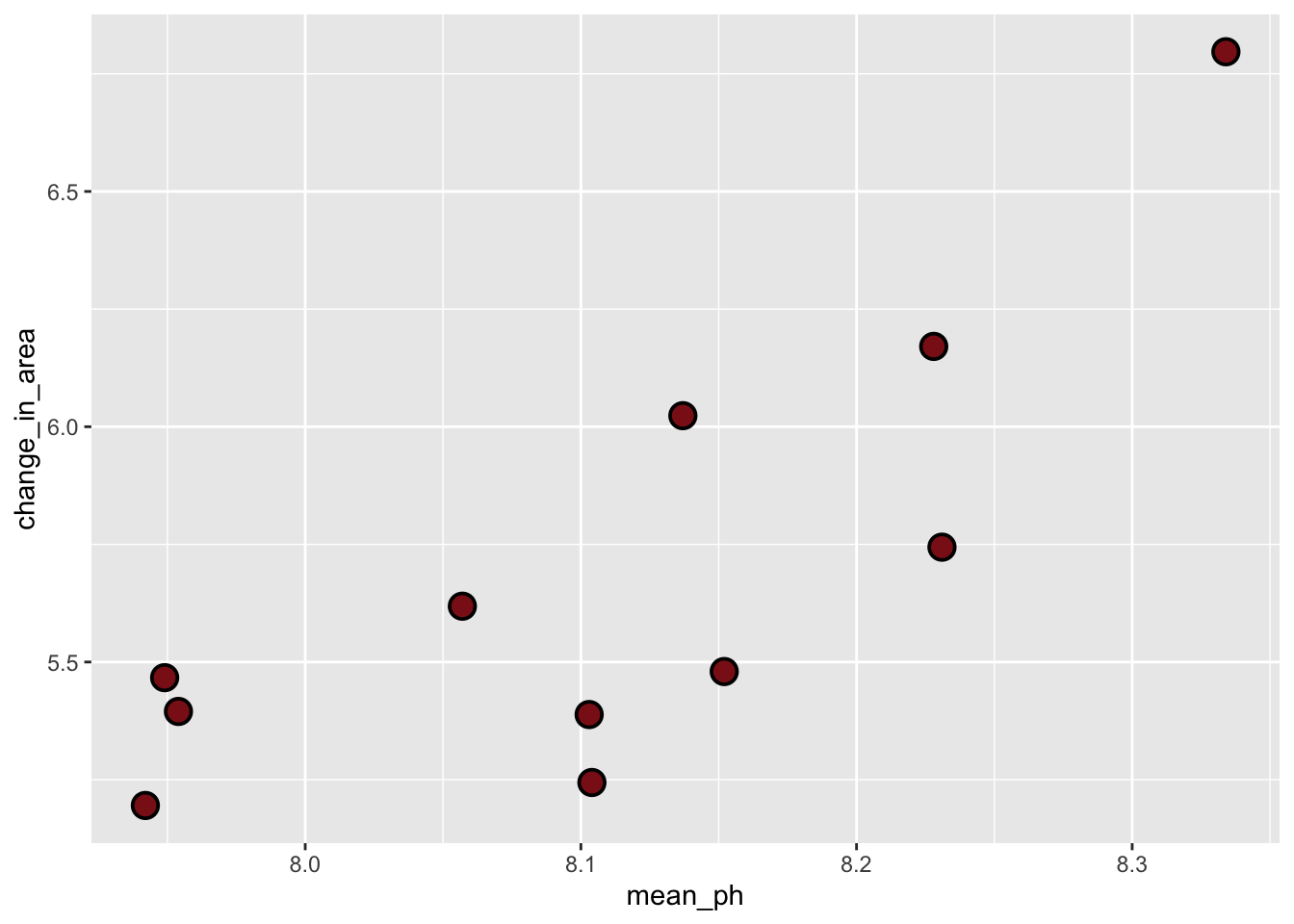
Where does the missing value come from?
The missing value came from Tank B1B
Don’t forget to turn your warnings off either in the individual code chunk or in the YAML at the top of the file.
d. Abalone model
Model fitting
abalone_model <- lm(
change_in_area ~ mean_ph, #
data = abalone_clean #
)Diagnostics
par(mfrow = c(2, 2)) #
plot(abalone_model) # 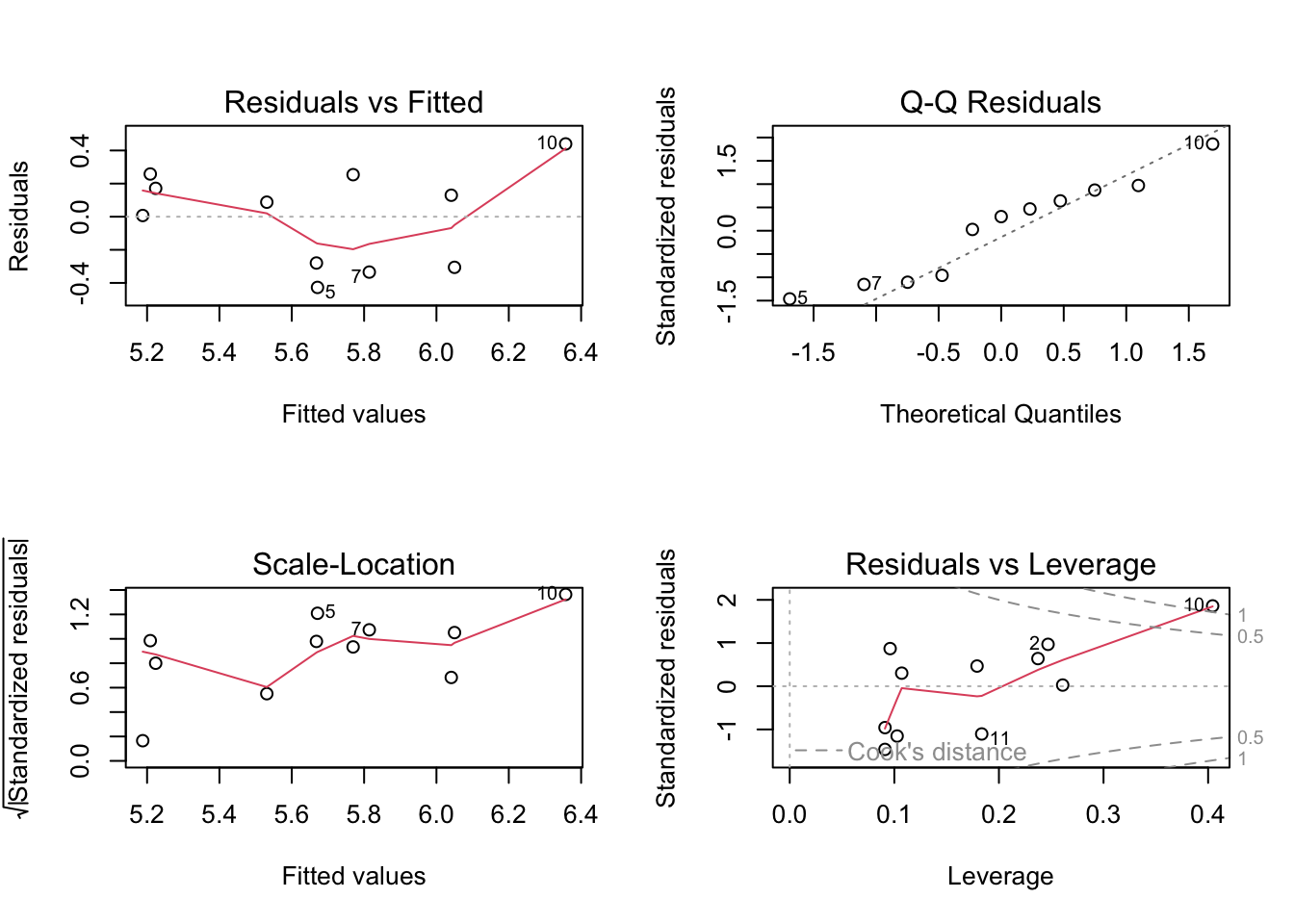
Residuals seem homoscedastic (consistent across values) based on Residuals vs. Fitted and Scale-Location plots. Residuals seem normally distributed based on the QQ plot, the points mostly follow the reference line. There were also no outliers that might influence model estimates or predictions based on Residuals vs. Leverage.
Model summary
# more information about the model
summary(abalone_model)
Call:
lm(formula = change_in_area ~ mean_ph, data = abalone_clean)
Residuals:
Min 1Q Median 3Q Max
-0.42714 -0.29282 0.08769 0.21258 0.43965
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -18.5010 6.1601 -3.003 0.01488 *
mean_ph 2.9827 0.7596 3.926 0.00348 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.3063 on 9 degrees of freedom
(1 observation deleted due to missingness)
Multiple R-squared: 0.6314, Adjusted R-squared: 0.5905
F-statistic: 15.42 on 1 and 9 DF, p-value: 0.003477What is the slope estimate (with standard error?)
With each invrease in pH, the model predicts an increase of 3.0 +/- in change in shell surface area per day (mm2 d-1)
What is the intercept estimate (with standard error?)
When pH = 0, the model predicts that abalone change in shell surface area per day is -18.5. (Note to self: this is not necessarily biologically realistic, but is something the model generates).
Generating predictions
# creating a new object called abalone_preds
abalone_preds <- ggpredict(
abalone_model, # model object
terms = "mean_ph" # predictor (in quotation marks)
)
# display the predictions
abalone_preds# Predicted values of change_in_area
mean_ph | Predicted | 95% CI
--------------------------------
7.94 | 5.19 | 4.83, 5.54
7.95 | 5.21 | 4.86, 5.55
8.06 | 5.53 | 5.30, 5.76
8.10 | 5.67 | 5.46, 5.88
8.10 | 5.67 | 5.46, 5.88
8.14 | 5.77 | 5.55, 5.98
8.15 | 5.81 | 5.59, 6.04
8.33 | 6.36 | 5.92, 6.80Look at the column names using colnames(abalone_preds) in the Console.
What are the column names?
x is predictor variable values for pH
predicted is the model predictions for change in shell surface area per day
std. error: (standard error describing precision in model predictions)
Conf. low and conf. high (lower and upper bounds of 95% CI around model predictions)
Look at the actual object (by clicking on it in Environment).
Look at the class of the object using class(abalone_preds) in the Console.
What is this type of object?
ggeffects output object and a data frame
View abalone_preds like a data frame.
Why generate model predictions?
This dataset does not include any actual observations of abalone growth at pH = 8.
However, the model can tell us a prediction for what abalone growth could be if pH = 8 based on the existing data.
Figure out what abalone growth the model would predict at pH = 8.
# finding the model prediction at a specific value
ggpredict(
abalone_model, # model object
terms = "mean_ph[8]" # predictor (in quotation marks) and predictor value in brackets
)# Predicted values of change_in_area
mean_ph | Predicted | 95% CI
--------------------------------
8 | 5.36 | 5.08, 5.64How would you interpret this?
At pH = 8, the model predicts that abalone change in shell surface area per day should be 5.36 mm2 day-1 (95% CI: [5.08, 5.64])
Visualizing model predictions and data
# base layer: ggplot
# using clean data frame
ggplot(data = abalone_clean,
aes(x = mean_ph,
y = change_in_area)) +
# first layer: points representing abalones
geom_point(size = 4,
stroke = 1,
fill = "firebrick4",
shape = 21) +
# second layer: ribbon representing confidence interval
# using predictions data frame
geom_ribbon(data = abalone_preds,
aes(x = x,
y = predicted,
ymin = conf.low,
ymax = conf.high),
alpha = 0.1) +
# third layer: line representing model predictions
# using predictions data frame
geom_line(data = abalone_preds,
aes(x = x,
y = predicted)) +
# axis labels
labs(x = "Mean pH",
y = expression("Change in shell area ("*mm^{2}~d^-1*")")) +
# theme
theme_minimal()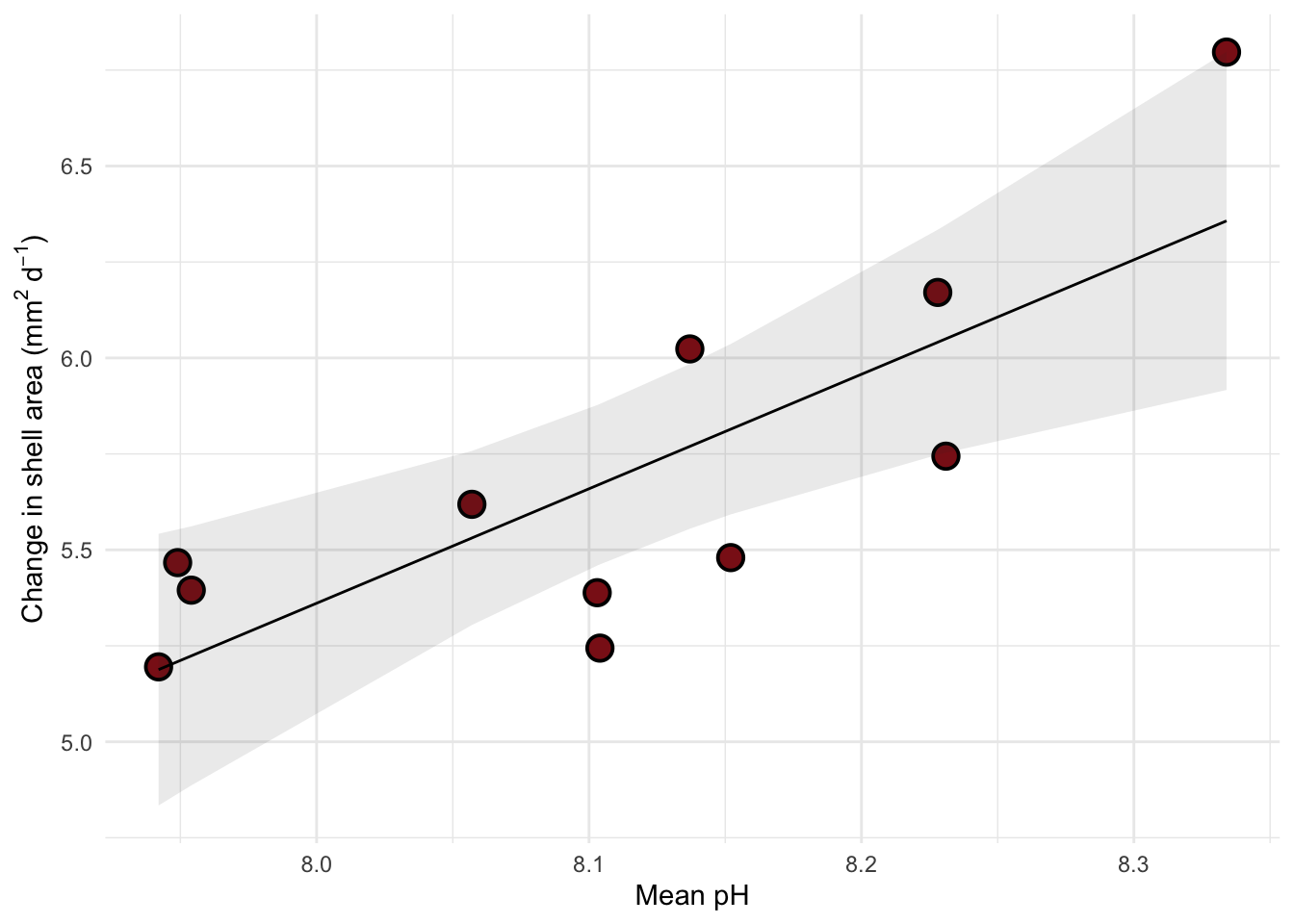
Creating a table with model coefficients, 95% confidence intervals, and more
Note: both these functions are from gtsummary.
tbl_regression(abalone_model,
# make sure the y-intercept estimate is shown
intercept = TRUE,
# changing labels in "Characteristic" column
label = list(`(Intercept)` = "Intercept",
mean_ph = "pH")) |>
# changing header text
modify_header(label = "**Variable**",
estimate = "**Estimate**") |>
# turning table into a flextable (makes things easier to render to word or PDF)
as_flex_table()Variable | Estimate | 95% CI | p-value |
|---|---|---|---|
Intercept | -19 | -32, -4.6 | 0.015 |
pH | 3.0 | 1.3, 4.7 | 0.003 |
Abbreviation: CI = Confidence Interval | |||
END OF ABALONE EXAMPLE